There are a number of utilities available at the BIC to view MINC volumes and other medical data sets from stand alone packages and from command line tools. In addition, MINC volumes can be easily converted and viewed online using a java applet.
BrainBrowser
 BrainBrowser is an open source JavaScript library exposing a set of web-based 3D visualization tools primarily targetting neuroimaging. Using open web-standard technologies, such as WebGL and HTML5, it allows for real-time manipulation and analysis of 3D imaging data through any modern web browser. The BrainBrowser Surface Viewer is a WebGL-based 3D viewer capable of displaying 3D surfaces in real-time and mapping various sorts of data to them. The BrainBrowser Volume Viewer is an HTML5 canvas-based viewer allowing slice-by-slice traversal of MINC volumetric data.
BrainBrowser is an open source JavaScript library exposing a set of web-based 3D visualization tools primarily targetting neuroimaging. Using open web-standard technologies, such as WebGL and HTML5, it allows for real-time manipulation and analysis of 3D imaging data through any modern web browser. The BrainBrowser Surface Viewer is a WebGL-based 3D viewer capable of displaying 3D surfaces in real-time and mapping various sorts of data to them. The BrainBrowser Volume Viewer is an HTML5 canvas-based viewer allowing slice-by-slice traversal of MINC volumetric data.
Standalone Visualization Software

IBIS (Intraoperative Brain Imaging System) Neuronav is an Image-Guided Neurosurgery planning and guidance software platform. It is meant to replicate the basic functionality of commercial image-guided systems such as Medtronic’s StealthStation while offering researchers an open system in which they can incorporate cutting-edge research modules for validation. On top of the basic features, Ibis provides a plugin system that allows developers to quickly implement new functionality.
Ibis is based on several open source technologies. The graphical user interface is built on Qt. Rendering of graphic window is done using the VTK framework augmented with some custom VTK classes. Additional data filtering is done with ITK and OpenCV. Ibis can render all kinds of medical images in both 2D slice views and 3D views. The 3D view can optionally contain volumetric rendering. In the OR, the software can be connected to an optical tracking system to register the patient to preoperative data. By attaching an optical tracking device to a video camera, Ibis is able to produce an augmented reality view of the operating field.
Publications:
Drouin, S., Kersten-Oertel, M., Chen, S., Collins, D.L. ‘’A Realistic Test and Development Environment for Mixed Reality in Neurosurgery’‘. Springer Lecture Notes In Computer Science, Vol. 7264, 13–23 (2012).
Mercier, L., Del Maestro, R.F., Petrecca, K., Kochanowska, A., Drouin, S., Yan, C., Janke, A. L., Chen, S., Collins, D.L. New prototype neuronavigation system based on preoperative imaging and intraoperative freehand ultrasound: system description and validation. Int J Comput Assist Radiol Surg 6(4), 507–522 (2011).
 Command-line Visualization Software
Command-line Visualization Software
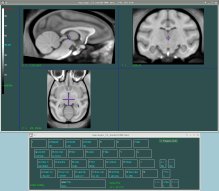
Display which is part of MINC tools, is the most complex of all of the visualization tools available. It’s primary uses are the display of multiple imaging modalities (PET, fMRI, etc.), as well as for manual segmentation of MRIs, aka “Brain Painting.”
 Register which is also part of MINC tools, can be used for superimposing two volumes as well as performing manual registrations
Register which is also part of MINC tools, can be used for superimposing two volumes as well as performing manual registrations
 xdisp is an image display tool for X window systems. It was written using basic Xlib calls (to make it easy to port to any X system) and has since had some (optional) EZwgl (Xlib based widgets) stuff added.
xdisp is an image display tool for X window systems. It was written using basic Xlib calls (to make it easy to port to any X system) and has since had some (optional) EZwgl (Xlib based widgets) stuff added.
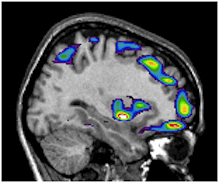
 Postf is a simple display program for medical image files, originally developed by Gabriel C. Léger at the McConnell Brain Imaging Centre.
Postf is a simple display program for medical image files, originally developed by Gabriel C. Léger at the McConnell Brain Imaging Centre.
 MNI ray_trace is not interactive; it is designed for the creation of static images from the command-line. Some of the advantages of this approach are reproducibility (i.e. it is much easier to specify the angle you want the picture taken from then to try to maneuver the volume into that position, in display) as well as cleaner image quality.
MNI ray_trace is not interactive; it is designed for the creation of static images from the command-line. Some of the advantages of this approach are reproducibility (i.e. it is much easier to specify the angle you want the picture taken from then to try to maneuver the volume into that position, in display) as well as cleaner image quality.
 JIV2 is a register-like program written in Java. It can be used to view more than one MINC volume related through an affine transform.
JIV2 is a register-like program written in Java. It can be used to view more than one MINC volume related through an affine transform.




